A World First! RORC Gene Defects Identified as the Cause of Primary Immunodeficiencies in Humans

We talked to Dr. Satoshi Okada, lecturer in Integrated Health Sciences at the Institute of Biomedical & Health Sciences.
(Interview: February 5, 2016, by PR Group)
Introduction
Lecturer Satoshi Okada is researching primary immunodeficiencies, mainly focusing on severe congenital neutropenia. Conducting joint research with Professor Jean-Laurent Casanova and others at The Rockefeller University in the U.S.A., Dr. Okada succeeded in identifying the gene responsible for a primary immunodeficiency that simultaneously causes chronic mucocutaneous candidiasis and Mendelian susceptibility to mycobacterial infections. The outcome of this study was published in July 2015 in the online edition of the U.S. scientific journal Science.
*The published paper can be found here.
It is hoped that this discovery will not only assist in understanding the pathology and prognosis of primary immunodeficiencies resulting from congenital genetic disorders, but also lead to the development of therapies for such conditions. We talked to Dr. Okada about the background to his discovery of the responsible gene and the prospects that it offers for the future.
Research focused on immunity
The immune system protects the body by fighting any bacteria and fungi that have gained access to it. Primary immunodeficiencies – the subject of Dr. Okada’s research – are disorders in which a person’s immune system fails to function properly from the time when they are born. If a person’s immune system does not function normally, they become more prone to infectious diseases and the aggravation thereof. Most congenital immune function abnormalities are thought to be caused by gene defects, so Dr. Okada is conducting research aimed at identifying the genes responsible for them and unraveling their pathology.

Dr. Okada says, “My research involves unraveling the mystery of genetic abnormalities.”
Humans are known to have around 22,000 genes, which function as a blueprint for life itself. Genetic disorders are closely linked to the onset of disease, so genetic analysis, in the form of genetic diagnosis, has recently begun to be used in patient diagnosis, as well. Previously, we had to analyze each individual target gene for genetic diagnosis. However, next-generation sequencers – devices capable of decoding a large volume of DNA base sequences quickly – began to be introduced in 2009 or thereabouts, making it easier to identify genetic mutations.
Breakthrough triggered by the unexpected discovery of a protein
While Dr. Okada was looking over a list of mutated genes in a patient with a mycobacterial infection, which a next-generation sequencer had analyzed, the gene RORC happened to catch his eye. RORC is a gene that encodes RORγt. A molecule that has recently become the focus of attention, RORγt is a protein that plays a role in the differentiation of Th17 cells, which are crucial to the elimination of bacteria that have entered the body, as well as fungi adhering to mucosal surfaces.
“Th17 cells produce interleukin-17. I knew to some extent from previous research that interleukin-17 is crucial in guarding against candida infections of the mucosa. Accordingly, I expected to find candidiasis in this patient with a RORγt defect. However, analysis of data from the patient, who had a severe mycobacterial infection, revealed an abnormality in the RORC gene, which really surprised me at first. It occurred to me that if I studied this patient, it might lead to a breakthrough in understanding the unknown functions of the RORC gene, RORγt, and Th17 cells,” says Dr. Okada, reflecting on the beginning of this study.
The first identification of germline bi-allelic mutations in the RORC gene in humans
As a result of analysis, Dr. Okada identified the RORC gene mutation found in the patient as a loss-of-function mutation that impaired the function of the gene. This was the first time in the world that a patient born lacking RORC gene function had been discovered. Through the discovery of this patient, the research team verified that the phenotypes* found in mice with defective Rorc genes (such as lack of peripheral lymph nodes, thymic hypoplasia, and reduced Th17 cells) were also found in humans.
(*) Phenotype: A trait encoded by a gene and expressed in an organism
Although Dr. Okada ascertained that the RORC gene mutation found in the patient was a loss-of-function mutation, he adds, “The relationship between the RORC gene and mycobacterial infection had remained a complete mystery in studies up to that point.” Further research revealed that patients with a loss-of-function mutation in the RORC gene suffer from impaired production of a cytokine called interferon-γ, which is required to guard against mycobacterial infection. Thus, the fact that the RORC gene is essential in guarding against not only candida infection, but also mycobacterial infection was discovered for the first time worldwide.

“These patients taught us the role of the RORC gene,” says Dr. Okada.
A turning point, just as he was ready to give up
Before going to study in the U.S.A., Dr. Okada set himself two goals: to discover a new gene responsible for causing a disease; and to publish an article detailing that discovery in a top journal such as Science. While he certainly managed to achieve those goals, the road to that achievement was not always an easy one.
From in vitro function tests, Dr. Okada was convinced that a mutation in the RORC gene caused the disease. “In the test cells, I forcibly expressed normal (wild-type) RORγ and RORγt, and (mutant) RORγ and RORγt with the mutations found in the patient, and then measured their capacity to bind with the target sequence. Whereas the wild-type protein bonded with the target sequence, bonding of the mutant protein to the target sequence was impaired. Looking at the results, I was certain that an abnormality in the RORC gene was the cause of the disease,” he recalls.
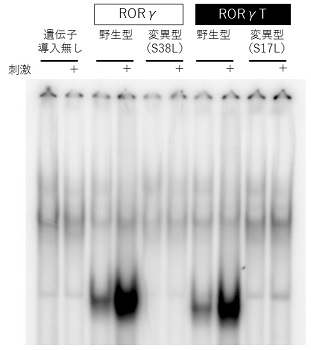
The test data from which Dr. Okada ascertained that the RORC gene mutation found in the patient was a loss-of-function mutation
However, he subsequently reached an impasse. “By the time I discovered the cause, the patient had already died. Accordingly, I was unable to use that patient’s blood to verify the cause of the disease. There was no way of checking whether the phenomenon that I’d observed in vitro could be found in the actual patient. I felt as though I’d reached the end of the road with this study,” says Dr. Okada, looking back.
Then, about a year after his discovery of the RORC gene defect, he reached a turning point. A second and third patient with RORC gene defects were found. Just like the first patient, these two patients were the offspring of marriages between close relatives. From three families, including the family members of the first patient in whom the genetic defect was discovered, Dr. Okada found a total of seven patients with the RORC gene mutation and began the process of analysis. This enabled him to verify his findings using the patients’ blood, so his research project got underway again.
Building “relationships of mutual reliance”
Having returned to Hiroshima University after completing his three-and-a-half-year period of study abroad, Dr. Okada continued his joint research with The Rockefeller University St. Giles Laboratory of Human Genetics of Infectious Diseases. As well as Hiroshima University and The Rockefeller University, this research project came to involve researchers from across the globe. Dr. Okada made progress with his research with the aid of numerous researchers, sometimes requesting their joint participation in the project because they had access to analytical technologies that he needed for this study.
The research “would not have been successful without their assistance,” says Dr. Okada. He maintained a firm stance of not saying no, if at all possible, when other researchers asked him to carry out tests for them, thereby cultivating “relationships of mutual reliance.” Apparently, he established this stance during his time studying abroad. “As well as U.S. researchers, there were researchers from various other countries at The Rockefeller University Laboratory of Human Genetics of Infectious Diseases where I was studying, including people from China, France, Germany, Colombia, and Iran, to name just a few. Naturally, there was some rivalry among us, but we didn’t try to hold each other back; rather, the atmosphere was one in which we encouraged each other to improve so that we could all grow as a result. It was an environment with really great teamwork,” says Dr. Okada. By helping each other out, the researchers were able to up the pace of the study and got close to unraveling the cause of the disease.
The importance of “clean” tests
When Dr. Okada carries out tests, there is one thing of which he is particularly conscious: conducting “clean” tests. Before carrying out tests, he first thinks carefully about the conditions and procedures, and then assembles the tests one by one. He mulls over the study and builds up a mental picture of the route to his goal before proceeding with the tests. “If I don’t get my thoughts in order before conducting the tests, I don’t get clean figures and test data,” he says. This advanced skill of conducting precise tests to generate clean results is one of Dr. Okada’s strengths. Although he says laughs that his English was not great during his time studying abroad, he says proudly that “my experimental skills were highly rated.” “My boss, Professor Casanova, praised me, saying, ‘Your test data are always clean, Satoshi.’ That experience gave me confidence, so I always tell the students I supervise to make sure that they conduct ‘clean’ tests,” says Dr. Okada.
Another thing that he always bears in mind is the importance of a control group against which test data can be compared. This approach is based on Prof. Casanova’s guidance. When conducting tests, he takes great care to ensure that he has both a positive control and a negative control, without fail. “One control alone, whether positive or negative, is not sufficient for evaluating test data. To provide a concrete explanation, I use the measurement of enzyme activity as an example. Enzyme activity in a healthy individual (positive control) was 100 and enzyme activity in the patient was 60. Compared with 100, 60 certainly is low, so you could say that the enzyme is defective. On the other hand, if there was an affected individual who was actually shown to be lacking the enzyme (negative control) and their enzyme activity level was 10, the evaluation of the patient with an enzyme activity level of 60 would change completely. To ensure that we can evaluate test results properly, it’s vital for us to have both a positive and a negative control in all tests,” Dr. Okada goes on to say.

Dr. Okada supervises a test conducted by graduate students.
Searching for the truth
No matter how carefully one carries out tests, only 10% of all data are suitable for use in an academic paper. Apparently, for various reasons, including technical problems with the tests, 90% of data are unable to be used in a paper. “It’s really tough when you can’t get acceptable test results. But I don’t think it’s right to obsess about getting results that match your hypothesis. I carry out tests over and over again, devising conditions and mulling over the interpretation of the results, so that the truth begins to emerge. I’m over the moon when I get data that I can use in a paper,” says Dr. Okada, with a smile. “I’m able to unravel with my own hands things that are not yet understood. The process of finding out new things is interesting in itself, and the moment you finally understand them is sheer bliss. When your intellectual curiosity is satisfied, research feels truly worthwhile. I want as many people as possible to experience the joy of discovering new things.”
Future prospects
There are many patients with primary immunodeficiencies whose responsible gene is still unknown. “There’s still ample potential for the discovery of new genes. I want to make a contribution to both medical science and patients through discoveries that will lead to breakthroughs in these diseases,” says Dr. Okada, enthusiastically talking about the future. Perhaps it will not be too long before we hear about another new discovery.

Afterword
Dr. Okada was kind enough to explain the tricky technical terms using words that were easy to understand. While I glimpsed the kindly face of this pediatrician, his zealous desire as a researcher to make discoveries that will benefit patients shone through during our conversation from beginning to end. I was shocked to discover that only 10% of all data are suitable for inclusion in an academic paper. I imagine that it takes a lot of hard work and effort, but I was impressed by Dr. Okada’s animated expression when he said, “There are so many fun and interesting things in research, you know!” After the interview, he was kind enough to show us around his laboratory, where graduate students chatted with him in a friendly way and seemed to be enjoying carrying out tests. Perhaps it is the good atmosphere in the laboratory that makes research more enjoyable. I hope that they will discover more new genes and unravel the causes of as many diseases as possible. (F)


 Home
Home

