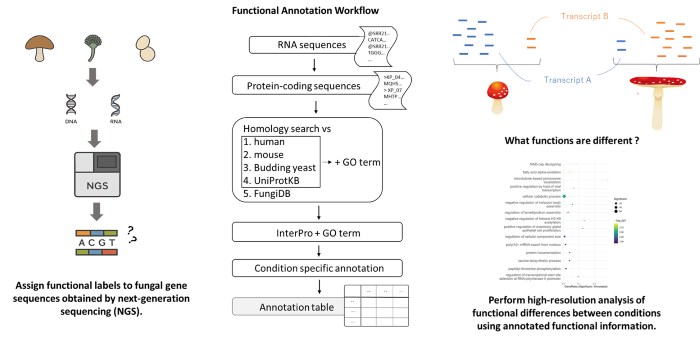
RNA and DNA sequences from fungal samples were generated by next-generation sequencing (NGS) and functionally annotated using the new workflow to construct a comprehensive annotation table. The annotated functional information enabled high-resolution functional enrichment analysis, revealing biologically meaningful differences between conditions. (Nagisa Morihara/Hiroshima University)
While RNA sequencing (RNA-seq) has become a standard tool for profiling which genes are active in an organism, determining the actual biological functions of those genes in fungi remains a significant technical challenge. Most existing software tools are designed to cover a broad range of species, often lacking the specificity required for the unique genetic landscape of fungi. Furthermore, many non-model fungal species lack the high-quality reference genomes that traditional analysis methods rely on.
To address these limitations, a research team led by Professor Hidemasa Bono at Hiroshima University’s Graduate School of Integrated Sciences for Life created a fungal-specific workflow that supports downstream functional analysis regardless of whether a reference genome is available.
“While RNA sequencing has become easier with next-generation sequencers, existing general-purpose tools fail to capture fungal-specific features, leaving a high proportion of genes functionally uncharacterized. This limitation has been a major obstacle for downstream analyses such as functional enrichment analysis and genome editing applications,” said Bono. “Our approach enables accurate functional interpretation of fungal transcriptomes and facilitates the identification of biologically meaningful genes that were previously overlooked.”
In the study, published in the Journal of Fungi on February 6, 2026, the researchers evaluated the workflow's performance by processing RNA-seq data from 57 samples of shiitake mushroom (Lentinula edodes strain H600) and 20 samples of Asian soybean rust (Phakopsora pachyrhizi) retrieved from public databases. By testing the system on these two biologically distinct fungi—one a widely cultivated edible mushroom and the other a devastating agricultural pathogen—the team demonstrated that the tool maintains high accuracy across diverse fungal lifestyles. The workflow successfully annotated over 96% of protein-coding transcripts, providing a much higher resolution of functional detection than existing generalized annotation tools.
Furthermore, the system proved its technical versatility by successfully processing data from two different types of sequencing data: standard short-read RNA-seq and full-length transcript sequencing (Iso-Seq). Instead of relying on a pre-existing reference genome, the researchers compared sequences against specialized fungal-specific databases and analyzed expression patterns.
“This improvement enables more comprehensive functional enrichment analysis that better reflects actual biological phenomena occurring during fungal development and pathogenesis,” said Bono.
The dual approach allows the workflow to pinpoint functionally important transcripts, even in non-model species, identifying high-priority targets for applications such as CRISPR-based genome editing or other biotechnological innovations. The researchers anticipate that this tool will significantly facilitate the discovery of functionally important transcripts in diverse fungal species, ultimately accelerating their translation into industrial and medical innovations.
The study was co-authored by Nagisa Morihara and Hidemasa Bono of Hiroshima University.

 Home
Home